Breast Cancer (Sklearn)
1. 1. Load Data
2. 2. Train Test Split
3. 3. KNeighborsClassifier
4. 4. Evaluation par Validation Croisée
5. 5. Recherche du meilleur Hyper-paramètre N_Neighbors
6. 7. Optimisation de plusieurs Hyper-paramètres
2. 2. Train Test Split
3. 3. KNeighborsClassifier
4. 4. Evaluation par Validation Croisée
5. 5. Recherche du meilleur Hyper-paramètre N_Neighbors
6. 7. Optimisation de plusieurs Hyper-paramètres
1. 1. Load Data
feature names
import seaborn as sns
from sklearn.datasets import load_breast_cancer
data = load_breast_cancer()
print(data.feature_names)
['mean radius' 'mean texture' 'mean perimeter' 'mean area' 'mean smoothness' 'mean compactness' 'mean concavity' 'mean concave points' 'mean symmetry' 'mean fractal dimension' 'radius error' 'texture error' 'perimeter error' 'area error' 'smoothness error' 'compactness error' 'concavity error' 'concave points error' 'symmetry error' 'fractal dimension error' 'worst radius' 'worst texture' 'worst perimeter' 'worst area' 'worst smoothness' 'worst compactness' 'worst concavity' 'worst concave points' 'worst symmetry' 'worst fractal dimension']
30 variables et 569 points
X = data.data
print(f'{X.shape=}')
(569, 30)
2 classes : '0' (212 lignes) et '1' (357 lignes)
y = data.target
print('y: ', np.unique(y, return_counts=True))
(array([0, 1]), array([212, 357]))
Catégories sur mean_radius et mean_texture
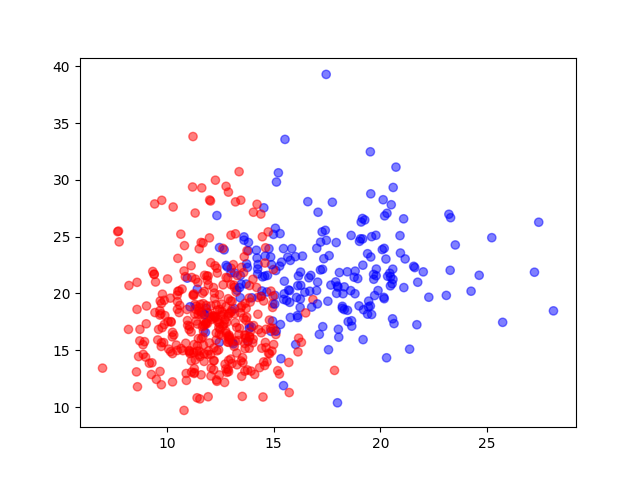
plt.scatter(X[:, 0], X[:, 1], c=y, alpha=0.5, cmap='bwr')

2. 2. Train Test Split
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)
Train set avec 455 points et Test set avec 114 points
print(f'{X_train.shape=}')
X_train.shape=(455, 30)
print(f'{X_test.shape=}')
X_test.shape=(114, 30)
print(f'{y_train.shape=}')
y_train.shape=(455,)
print(f'{y_test.shape=}')
y_test.shape=(114,)
3. 3. KNeighborsClassifier
from sklearn.neighbors import KNeighborsClassifier
model = KNeighborsClassifier(n_neighbors=1)
model.fit(X_train, y_train)
train_predictions = model.predict(X_train)
test_prediction = model.predict(X_test)
train_accuracy = accuracy_score(y_train, train_predictions)
test_accuracy = accuracy_score(y_test, test_prediction)
Train Accuracy avec n_neighbors=1
train_accuracy = accuracy_score(y_train, train_predictions)
1.0
Test Accuracy avec n_neighbors=1
test_accuracy = accuracy_score(y_test, test_prediction)
0.9122807017543859
4. 4. Evaluation par Validation Croisée
from sklearn.model_selection import cross_val_score
model = KNeighborsClassifier(n_neighbors=1)
cv_scores = cross_val_score(
estimator=model,
X=X_train,
y=y_train,
cv=5,
scoring='accuracy'
)
Scores de validation croisée
5 cv_scores (folds, splits)
[0.94505495 0.89010989 0.9010989 0.94505495 0.92307692]
Moyenne des scores de validation croisée
cv_scores.mean()
0.9208791208791209
5. 5. Recherche du meilleur Hyper-paramètre N_Neighbors
Validation croisée sur hyper-paramètre n_neighbors
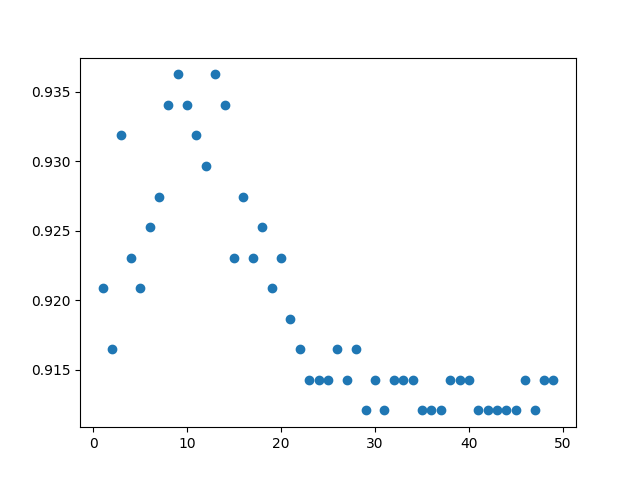
k_scores = {}
for k in range(1, 50):
model = KNeighborsClassifier(n_neighbors=k)
cv_scores = cross_val_score(
estimator=model, X=X_train, y=y_train, cv=5, scoring='accuracy'
)
k_scores[k] = cv_scores.mean()
plt.scatter(k_scores.keys(), k_scores.values())

6. 7. Optimisation de plusieurs Hyper-paramètres
param_grid = {
'n_neighbors': range(1, 20),
'metric': ['minkowski', 'cosine', 'euclidean']
}
grid_search = GridSearchCV(estimator=model, param_grid=param_grid, cv=5, scoring='accuracy')
grid_search.fit(X_train, y_train)
print(grid_search.cv_results_)
Contenu du dictionnaire
grid_search.cv_results_.keys
dict_keys(['mean_fit_time', 'std_fit_time', 'mean_score_time', 'std_score_time', 'param_metric', 'param_n_neighbors', 'params', 'split0_test_score', 'split1_test_score', 'split2_test_score', 'split3_test_score', 'split4_test_score', 'mean_test_score', 'std_test_score', 'rank_test_score'])
Paramètres testés
params = grid_search.cv_results_['params']
[{'metric': 'minkowski', 'n_neighbors': 1}, {'metric': 'minkowski', 'n_neighbors': 2}, {'metric': 'minkowski', 'n_neighbors': 3}, {'metric': 'minkowski', 'n_neighbors': 4}, {'metric': 'minkowski', 'n_neighbors': 5}, {'metric': 'minkowski', 'n_neighbors': 6}, {'metric': 'minkowski', 'n_neighbors': 7}, {'metric': 'minkowski', 'n_neighbors': 8}, {'metric': 'minkowski', 'n_neighbors': 9}, {'metric': 'minkowski', 'n_neighbors': 10}, {'metric': 'minkowski', 'n_neighbors': 11}, {'metric': 'minkowski', 'n_neighbors': 12}, {'metric': 'minkowski', 'n_neighbors': 13}, {'metric': 'minkowski', 'n_neighbors': 14}, {'metric': 'minkowski', 'n_neighbors': 15}, {'metric': 'minkowski', 'n_neighbors': 16}, {'metric': 'minkowski', 'n_neighbors': 17}, {'metric': 'minkowski', 'n_neighbors': 18}, {'metric': 'minkowski', 'n_neighbors': 19}, {'metric': 'cosine', 'n_neighbors': 1}, {'metric': 'cosine', 'n_neighbors': 2}, {'metric': 'cosine', 'n_neighbors': 3}, {'metric': 'cosine', 'n_neighbors': 4}, {'metric': 'cosine', 'n_neighbors': 5}, {'metric': 'cosine', 'n_neighbors': 6}, {'metric': 'cosine', 'n_neighbors': 7}, {'metric': 'cosine', 'n_neighbors': 8}, {'metric': 'cosine', 'n_neighbors': 9}, {'metric': 'cosine', 'n_neighbors': 10}, {'metric': 'cosine', 'n_neighbors': 11}, {'metric': 'cosine', 'n_neighbors': 12}, {'metric': 'cosine', 'n_neighbors': 13}, {'metric': 'cosine', 'n_neighbors': 14}, {'metric': 'cosine', 'n_neighbors': 15}, {'metric': 'cosine', 'n_neighbors': 16}, {'metric': 'cosine', 'n_neighbors': 17}, {'metric': 'cosine', 'n_neighbors': 18}, {'metric': 'cosine', 'n_neighbors': 19}, {'metric': 'euclidean', 'n_neighbors': 1}, {'metric': 'euclidean', 'n_neighbors': 2}, {'metric': 'euclidean', 'n_neighbors': 3}, {'metric': 'euclidean', 'n_neighbors': 4}, {'metric': 'euclidean', 'n_neighbors': 5}, {'metric': 'euclidean', 'n_neighbors': 6}, {'metric': 'euclidean', 'n_neighbors': 7}, {'metric': 'euclidean', 'n_neighbors': 8}, {'metric': 'euclidean', 'n_neighbors': 9}, {'metric': 'euclidean', 'n_neighbors': 10}, {'metric': 'euclidean', 'n_neighbors': 11}, {'metric': 'euclidean', 'n_neighbors': 12}, {'metric': 'euclidean', 'n_neighbors': 13}, {'metric': 'euclidean', 'n_neighbors': 14}, {'metric': 'euclidean', 'n_neighbors': 15}, {'metric': 'euclidean', 'n_neighbors': 16}, {'metric': 'euclidean', 'n_neighbors': 17}, {'metric': 'euclidean', 'n_neighbors': 18}, {'metric': 'euclidean', 'n_neighbors': 19}]
Scores obtenus
scores = grid_search.cv_results_['mean_test_score']
[0.92087912 0.91648352 0.93186813 0.92307692 0.92087912 0.92527473 0.92747253 0.93406593 0.93626374 0.93406593 0.93186813 0.92967033 0.93626374 0.93406593 0.92307692 0.92747253 0.92307692 0.92527473 0.92087912 0.9010989 0.9010989 0.92087912 0.91648352 0.91868132 0.91648352 0.92087912 0.91868132 0.92087912 0.92527473 0.92087912 0.91868132 0.92527473 0.92087912 0.92307692 0.91868132 0.91868132 0.91428571 0.91428571 0.92087912 0.91648352 0.93186813 0.92307692 0.92087912 0.92527473 0.92747253 0.93406593 0.93626374 0.93406593 0.93186813 0.92967033 0.93626374 0.93406593 0.92307692 0.92747253 0.92307692 0.92527473 0.92087912]
grid_search_results = pd.DataFrame(params)
grid_search_results['score'] = scores
metric n_neighbors score 0 minkowski 1 0.920879 1 minkowski 2 0.916484 2 minkowski 3 0.931868 3 minkowski 4 0.923077 4 minkowski 5 0.920879 5 minkowski 6 0.925275 6 minkowski 7 0.927473 7 minkowski 8 0.934066 8 minkowski 9 0.936264 9 minkowski 10 0.934066 10 minkowski 11 0.931868 11 minkowski 12 0.929670 12 minkowski 13 0.936264 13 minkowski 14 0.934066 14 minkowski 15 0.923077 15 minkowski 16 0.927473 16 minkowski 17 0.923077 17 minkowski 18 0.925275 18 minkowski 19 0.920879 19 cosine 1 0.901099 20 cosine 2 0.901099 21 cosine 3 0.920879 22 cosine 4 0.916484 23 cosine 5 0.918681 24 cosine 6 0.916484 25 cosine 7 0.920879 26 cosine 8 0.918681 27 cosine 9 0.920879 28 cosine 10 0.925275 29 cosine 11 0.920879 30 cosine 12 0.918681 31 cosine 13 0.925275 32 cosine 14 0.920879 33 cosine 15 0.923077 34 cosine 16 0.918681 35 cosine 17 0.918681 36 cosine 18 0.914286 37 cosine 19 0.914286 38 euclidean 1 0.920879 39 euclidean 2 0.916484 40 euclidean 3 0.931868 41 euclidean 4 0.923077 42 euclidean 5 0.920879 43 euclidean 6 0.925275 44 euclidean 7 0.927473 45 euclidean 8 0.934066 46 euclidean 9 0.936264 47 euclidean 10 0.934066 48 euclidean 11 0.931868 49 euclidean 12 0.929670 50 euclidean 13 0.936264 51 euclidean 14 0.934066 52 euclidean 15 0.923077 53 euclidean 16 0.927473 54 euclidean 17 0.923077 55 euclidean 18 0.925275 56 euclidean 19 0.920879
GridSearchCV Heatmap for KNN
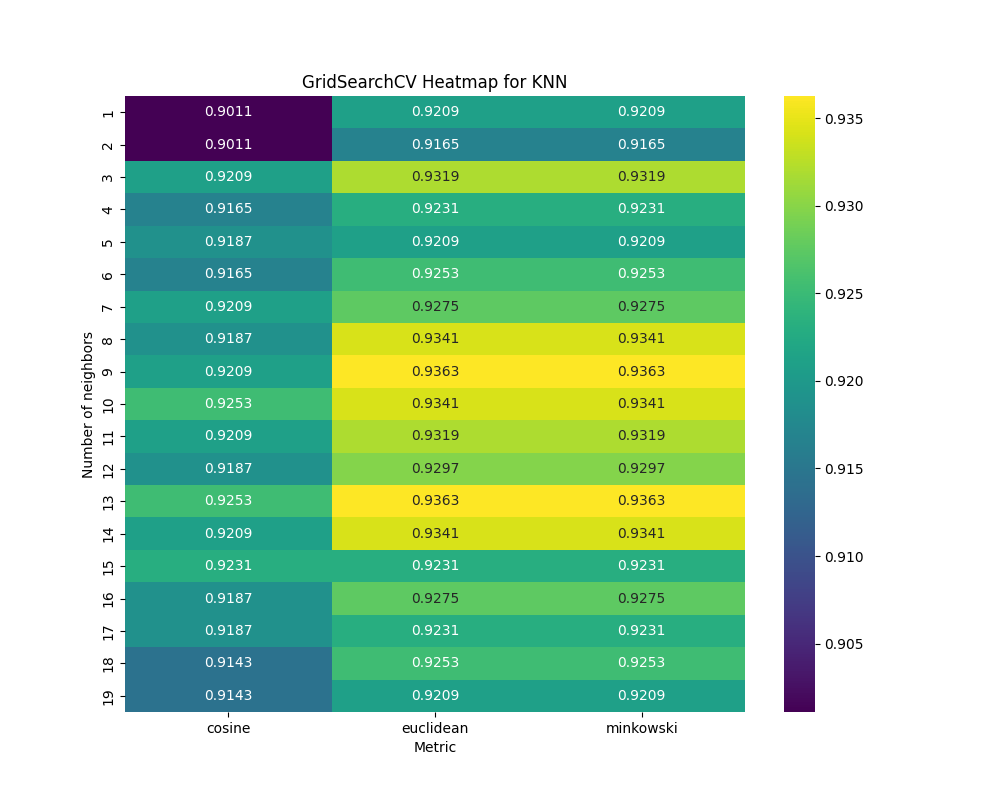
grid_search_heatmap = grid_search_results.pivot(index='n_neighbors', columns='metric', values='score')
plt.figure(figsize=(10,8))
sns.heatmap(grid_search_heatmap, annot=True, fmt='.4f', cmap='viridis')
plt.title('GridSearchCV Heatmap for KNN')
plt.xlabel('Metric')
plt.ylabel('Number of neighbors')

Meilleurs paramètres
grid_search.best_params_
{'metric': 'minkowski', 'n_neighbors': 9}
Meilleur score
grid_search.best_score_
0.9362637362637363
Accuracy sur le jeu de test
best_model = grid_search.best_estimator_
y_pred = best_model.predict(X_test)
test_accuracy = accuracy_score(y_test, y_pred)
0.9649122807017544